
MaCPepDB: A Database to Quickly Access All Tryptic Peptides of the UniProtKB | Journal of Proteome Research

The IDAES process modeling framework and model library—Flexibility for process simulation and optimization - Lee - 2021 - Journal of Advanced Manufacturing and Processing - Wiley Online Library

The Key Role of Metal Adducts in the Differentiation of Phosphopeptide from Sulfopeptide Sequences by High-Resolution Mass Spectrometry | Analytical Chemistry

Ursgal, Universal Python Module Combining Common Bottom-Up Proteomics Tools for Large-Scale Analysis | Journal of Proteome Research

LC–MS peak assignment based on unanimous selection by six machine learning algorithms | Scientific Reports
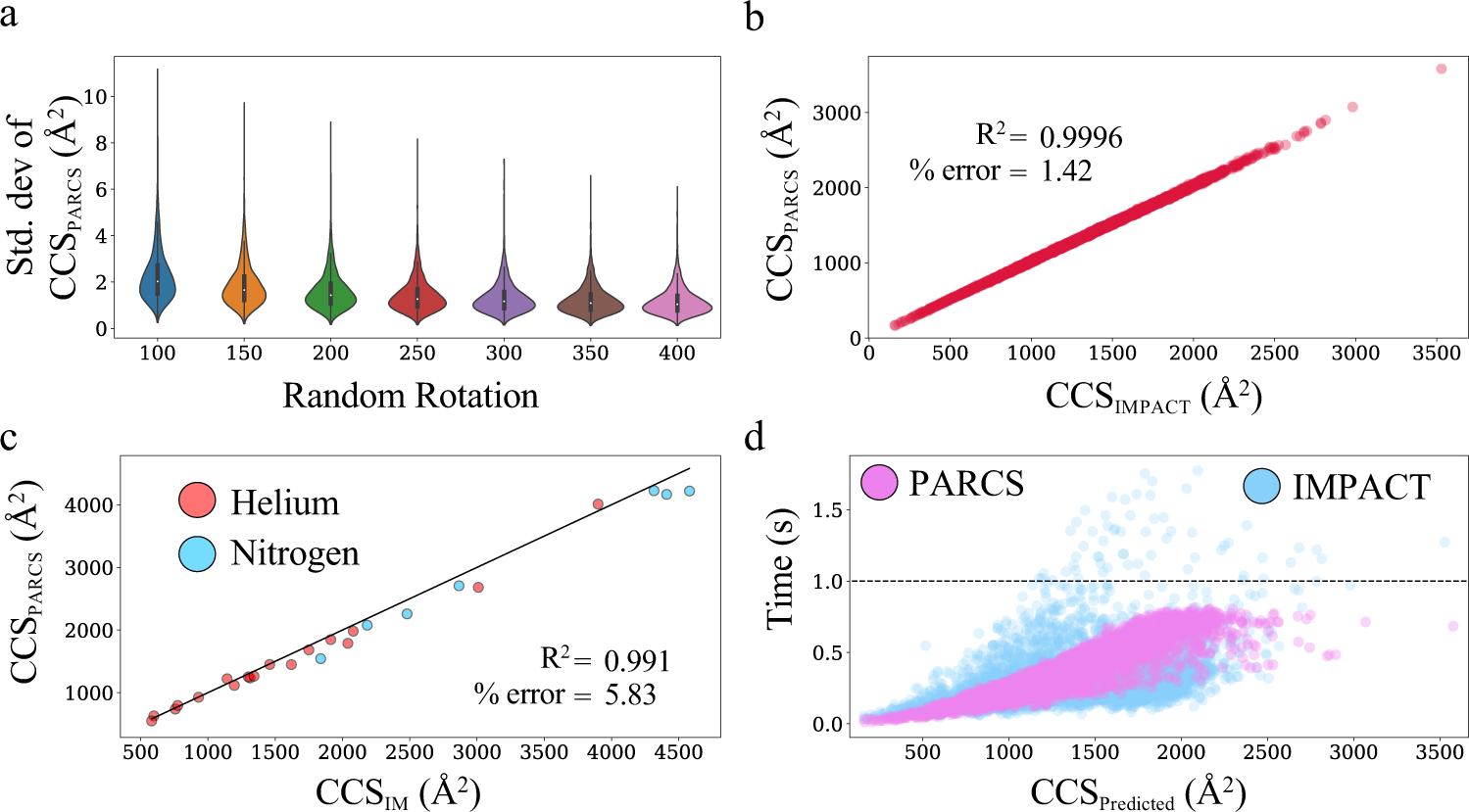
Protein shape sampled by ion mobility mass spectrometry consistently improves protein structure prediction | Nature Communications
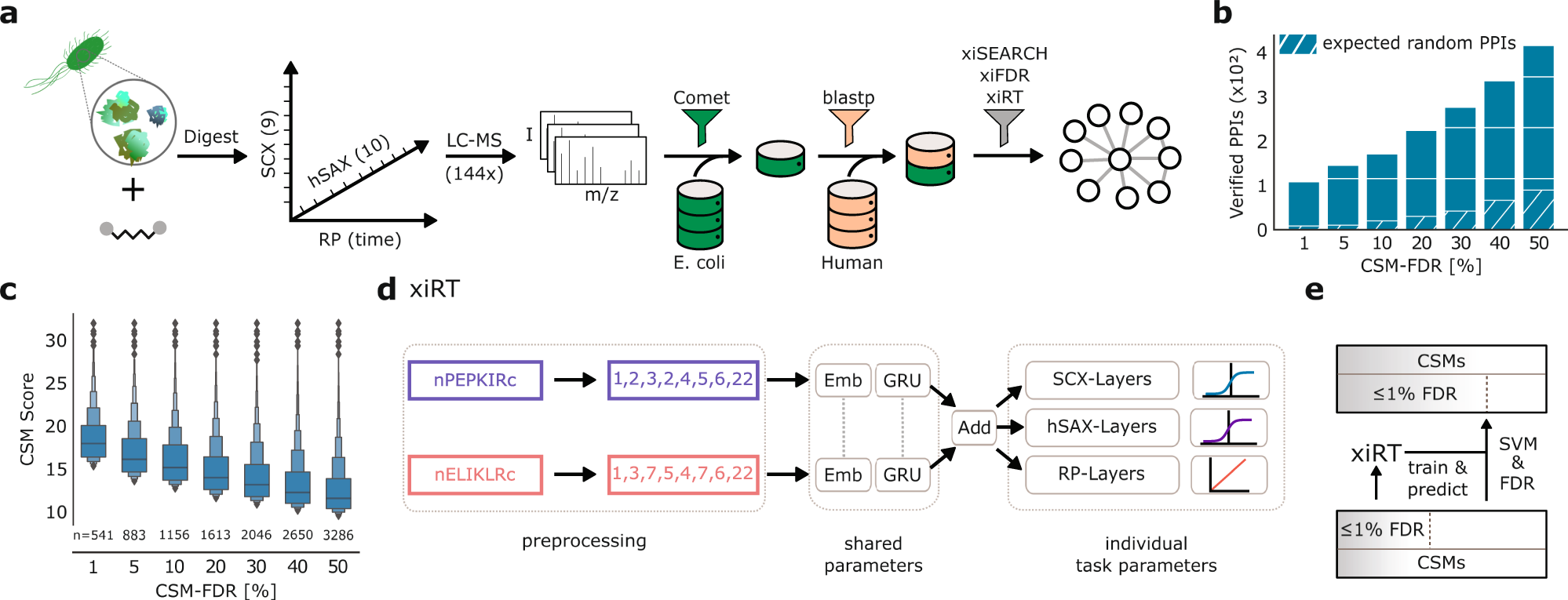
Retention time prediction using neural networks increases identifications in crosslinking mass spectrometry | Nature Communications

A calculated isotope distribution. The mass spectrum of a peptide or... | Download Scientific Diagram

pytc: Open-Source Python Software for Global Analyses of Isothermal Titration Calorimetry Data | Biochemistry








![Using genetic programming to predict and optimize protein function [PeerJ] Using genetic programming to predict and optimize protein function [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2022/pchem-24/1/fig-2-full.png)